Introduction
We use our own custom simulation code to conduct our molecular dynamics research. It shares the same general features as most downloadable (or "off-the-shelf") MD programs, but it is uniquely optimized for the sort of work we do in materials science. It lacks many of the complications that are necessary for the MD simulations conducted in inorganic chemistry or biological sciences, and as such, it tends to be much faster. A trade-off to this is that our code lacks many luxuries of big software packages. There is no graphical interface, and configuring simulations requires editing text files by hand.
General Program Flow
When executed, our simulation code (typically called something like mdv.x)
reads in a specific set of files as input, carries out the simulation based on
the data in those input files, then produces a set of output files. The most
basic MD run operates like this:
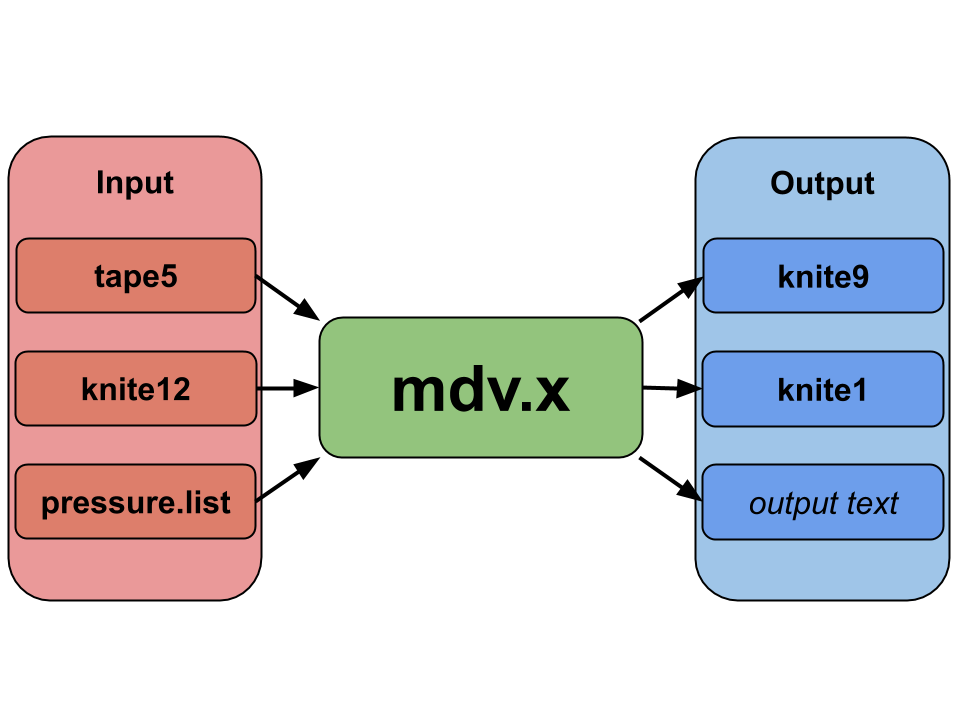
The MD code requires a files called tape5, pressure.list, and knite12 in
order to run, and without these three files, the simulation will instantly
crash. Once the code finds these files, it will run the simulation and virtually
always produce a file called knite1 and knite9. The code also prints a lot
of diagnostic output to the screen; most of the time, we redirect this output
to a third file with an arbitrary name. If these output files already exist
and another simulation is run, the old files will be overwritten without
warning, so be careful!
These filenames are hard-coded (e.g., if you have a file called tape5.6000
instead of tape5, the simulation will immediately crash because it only looks
for a file called tape5) and have unintuitive names, so here is a brief
explanation of what each one is:
knite12is a binary file that defines the starting configuration of the system to be simulated. This file describes every atom's position, velocity, acceleration, element, the size of the simulation box, etc.tape5is a text file that describes virtually all of the conditions of the simulation. This file contains thermodynamic and kinetic parameters such as the temperature at which you want to run and how long you want to simulation to last. It also contains all of the computational parameters such as the time step, interaction cutoff distance, and every empirical parameter for the interatomic potential (the model describing how the atoms interact) for all interactions.pressure.listis a very short text file that defines how the system's pressure is handled. This file is what you edit if you want to change from a constant volume simulation to a constant pressure simulation.mdv.xis the executable binary that does all the simulating. Different versions have different names (e.g.,mdvp.x,mdvgg.x, ormdvgg4040.x), but they are all based on the same set of source code.knite9is a binary file that defines the ending configuration of a simulation. It has the same format as theknite12file, and in factknite12files are often just the knite9 files of past simulations.- the
outputis printed to the screen by default but is often dumped into a text file so it can be reviewed even if nobody is sitting in front of the terminal. Its name varies, but it contains a log of the entire simulation as well as diagnostic information such as average and instantaneous pressures, temperatures, volumes, densities, energies, etc. knite1is a collection of configurations saved throughout the simulation. You can think of it as a series of knite9 files concatenated into a single file, and this is what allows us to analyze the simulation after it happened. If you think ofknite9is a single photo snapshot of the simulation,knite1would be the whole movie. And in fact you can generate movies of simulations using the data in this file.
Next page: Editing tape5